Accurate proteome-wide missense variant effect prediction with AlphaMissense
Por um escritor misterioso
Descrição
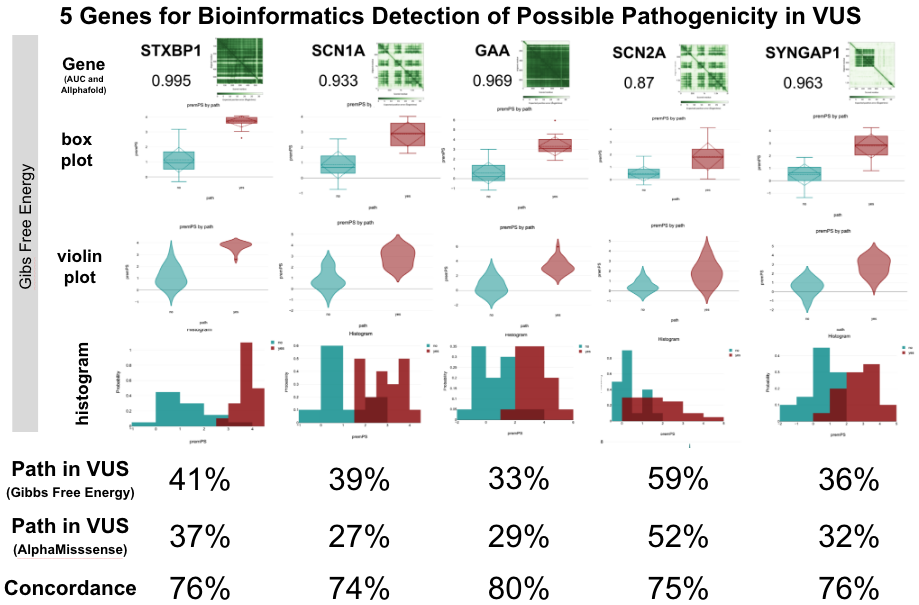
How Gene Data Sheds Light on Epilipesy

Google DeepMind Introduces AI Tool to Predict Harmful Genetic Mutations

Accurate prediction of protein tertiary structural changes induced by single-site mutations with equivariant graph neural networks

Predicting functional effect of missense variants using graph attention neural networks
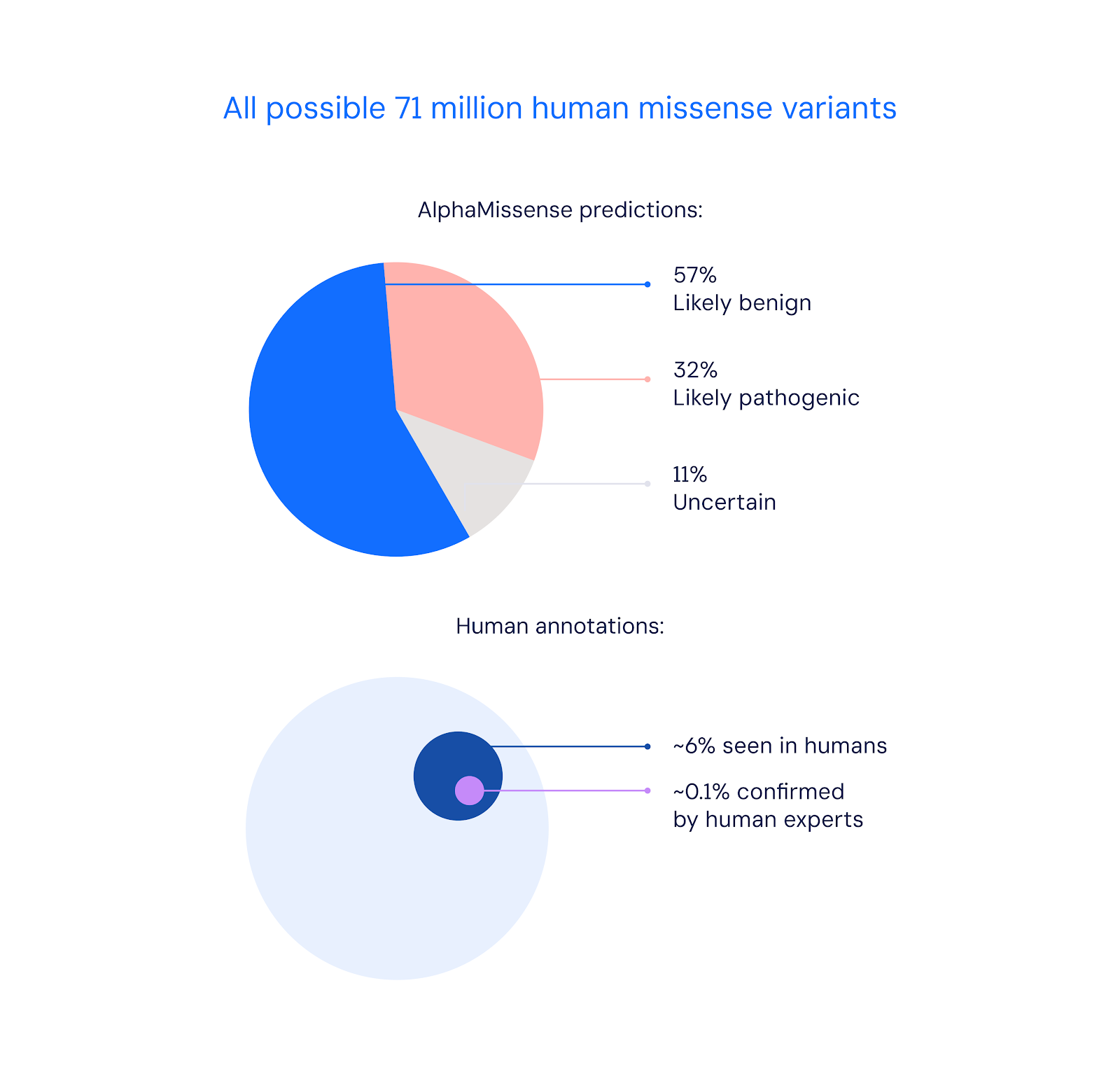
Meet AlphaMissense: Google's New AI Tool that Can Identify Protein Mutations Likely to Cause Disease

Can Predicted Protein 3D Structures Provide Reliable Insights into whether Missense Variants Are Disease Associated?
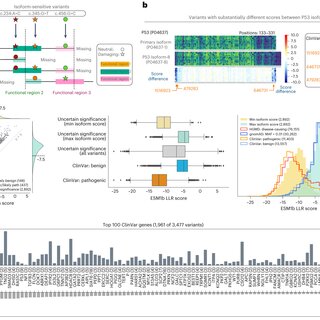
ESM1b predictions in clinically relevant genes depend on the isoform

PDF] Improved pathogenicity prediction for rare human missense variants
Google DeepMind announces AI ``AlphaMissense'', which may help predict which genetic mutations are harmful and identify the cause of genetic diseases - GIGAZINE
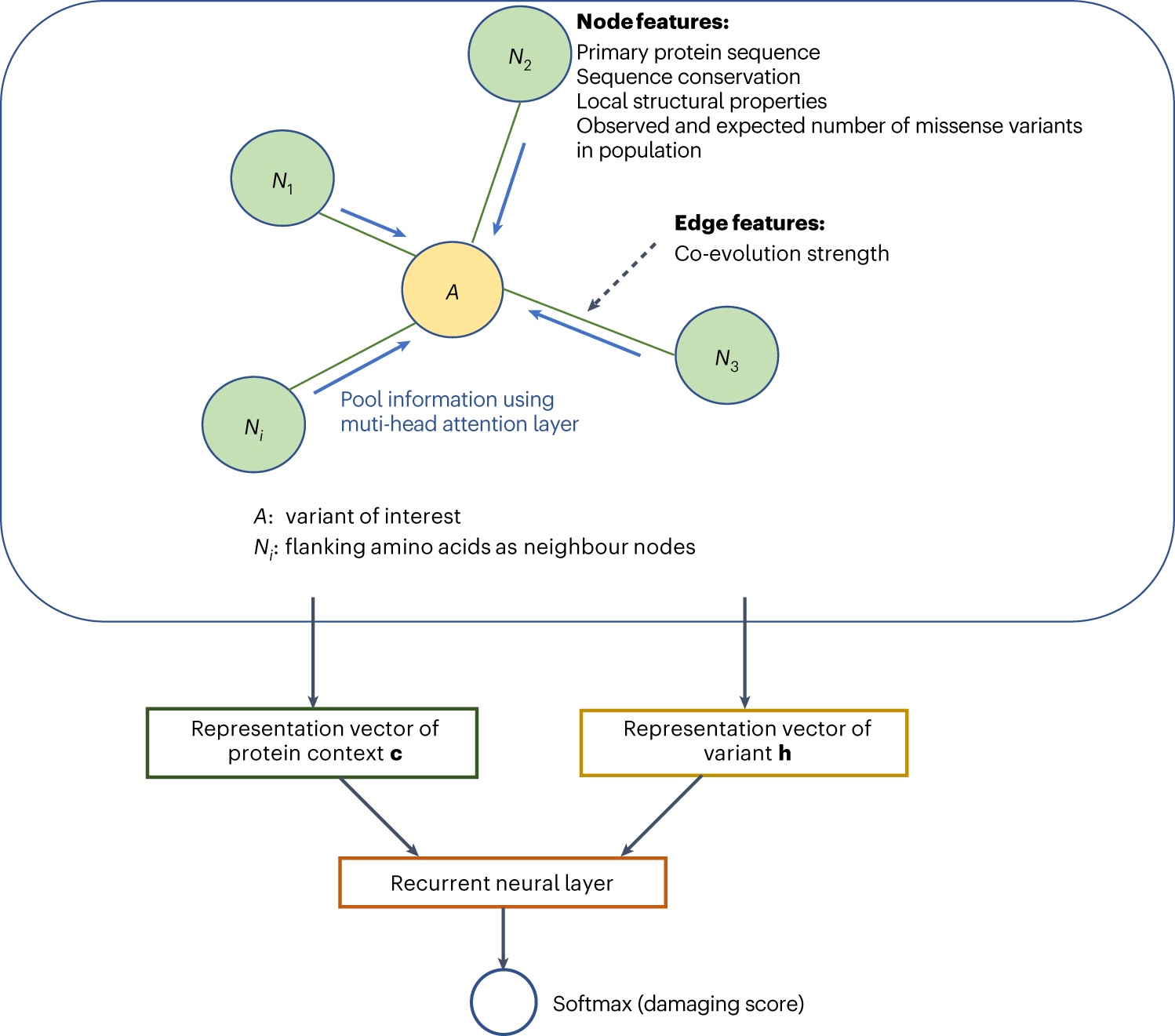
Predicting functional effect of missense variants using graph attention neural networks
GitHub - StephenStaklinski/alphamissense_asns: Snakemake pipeline for visualizing AlphaMissense pathogenicity score by UniProtID. Analysis of Asparagine Synthetase predictions.

Can Predicted Protein 3D Structures Provide Reliable Insights into whether Missense Variants Are Disease Associated?
de
por adulto (o preço varia de acordo com o tamanho do grupo)







